In the previous entries to this series I talked about computing power for an interaction analysis (Part 1), interpreting interaction effect-sizes (Part 2), and what sample size is needed for an interaction (Part 3). Here I’m going to talk about an issue that is near and dear to my heart – control variables!
“What?” you ask, “aren’t interaction control variables the same as any other regression?” The answer is simply…NO!
Let’s say you have a regression – Y ~ X + C. ‘C‘ here are my ‘control’ variables – variables that index potential effects which we’d like to account for. If we want to test an interaction, aka ‘moderation’, our regression might look like – Y ~ X*M + X + M + C. ‘X*M’ is the interaction that we are interested in testing. But we aren’t done! We haven’t adjusted for all possible effects of our control variables. Intuitively, since our hypothesis is an interaction, we need to control for potential alternative interactions. Our final regression equation should look like – Y ~ X*M + X + M + C*X + C*M + C.
I was first made aware of this issue by an excellent article from Dr. Matthew Keller on statistical controls for gene x environment analyses. He cites an article by Hull, Tedlie, and Hehn from 1992 that identified this problem (it’s the oldest article I know of that talks about it). There is also a great paper by Yzerbyt, Muller, and Judd from 2004, which in my opinion presents the best explanation of the underlying factors that drive the necessity of these additional controls.
Really, you should go read that paper by Yzerbyt, Muller, and Judd. To briefly summarize, there are two situations where a regression model that does not include covariate x predictor interaction terms will have increased false-positives: (1) when the covariate and predictor are correlated, and (2) when the covariate x predictor interaction is itself associated with the the dependent variable Y. Here I reproduce their Figure 1, showing these effects:
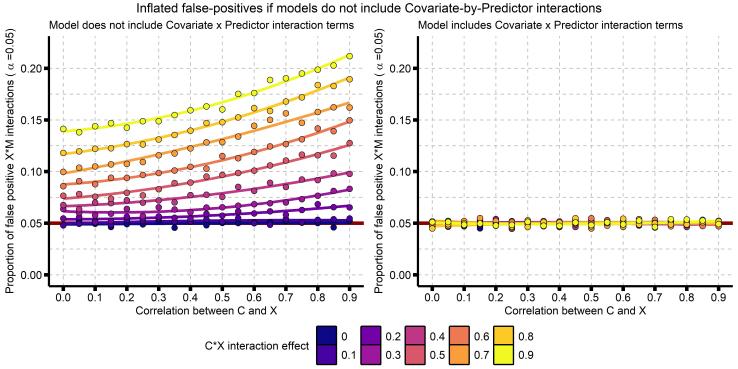
This plot makes it clear that the probability of a false-positive (a type-I error; aka detecting a significant effect when the true effect is 0) increases as the correlation between our covariate and predictor increases, and as the effect of our covariate x predictor interaction increases. Is this a problem? I really think it is. Even a false-positive-rate of 6% could be a huge issue for any individual finding, and at the level of a field or body of knowledge, could result in a highly biased literature. (Note this isn’t field-specific, this applies to any interaction analysis, so I’m not just talking about psychology or neuroscience here). I don’t have good data on how big of a problem this is, but I have read and reviewed so many papers that don’t include these additional covariates.
The extremely simple solution is to include all covariate-by-predictor interactions in your model. Of course, one down-side to this solution is that this will triple the number of covariates in your model – if you had 5 covariates, you will now have 15! This might be a problem if you have few degrees of freedom, though honestly, if you have so few df that an extra 10 variables makes a large difference, you probably shouldn’t be testing an interaction in the first place. You might wonder, what if you were to test each covariate for a potential source of bias, and only include the additional terms if they’re significant? It’s pretty clear why this won’t work, because (1) first you’ll have to correct for every statistical test (for instance 20 in my example of 5 initial covariates – the correlation between C and X/M, plus the effect of the C*X and C*M interactions), at which point (2) Figure 2 makes it clear that there is a clear possibility that you will be underpowered to detect an effect that will still be biasing your results (Yzerbyt, Muller, and Judd also make this point). So there’s no guarantee that you’ll be able to include only potential sources of biases. The only solution is to include all the interaction covariates.
Now, how to generate the covariates for your analyses? As far as I’m aware, plug-and-play statistical software like SPSS and GraphPad do not include an easy way to generate these variables on-the-fly, though you certainly could manually generate them one-by-one (or if you know how to script in SPSS it should also be quite fast). However, one thing that I’ve come to really appreciate about R is that these interaction covariates can be specified directly in in your regression model. For instance, instead of:
lm(Y~ X:M + X + M + C)
You would write:
lm(Y~ X:M + X + M + C + C:(X+M))
It really is that simple!
EDIT: I got a question about this, so for an ANCOVA analysis this would look like:
I would run a 2-way ANOVA as:
mod<-lm(y~1)
mod1<-lm(y~X:M + X + M)
anova(mod, mod1)
With a covariate C, a 2-way ANCOVA would be:
mod<-lm(y~C + C:(X+M)) mod1<-lm(y~X:M + X + M +C + (X:M)) anova(mod,mod1)
Adding Z for a 3-way ANCOVA:
mod<-lm(y~ C + C:(X:M:Z)+C:((X:M)+(Z:M)+(Z:X)) ) mod1<-lm(y~Z:X:M +X:M+Z:M+Z:X+ Z+ X + M +C + C:(X+M+Z)+C:((X:M)+(Z:M)+(Z:X))) anova(mod,mod1)


[…] that interactions need large sample sizes because we’re interested in small effects. In part 4 I then go on to discuss what covariates are needed when conducting an interaction […]
LikeLike
[…] little bit of extra intuition can greatly simplify power analyses for interactions. Finally, the fourth post, I go on to discuss which covariates are needed when conducting a test of an […]
LikeLike
[…] interaction effect-sizes in terms of their ‘simple slopes’. In the next post, Part 4, I’ll cover which covariates are needed when testing an […]
LikeLike